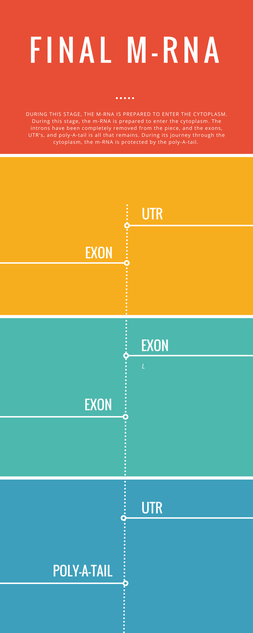
Infographic
Video Reflection
Throughout our video many good ideas were conveyed, but every model has mistakes. Sometimes in the video we would say the wrong word from the script, or label something incorrectly like centrosome instead of centromere or meiosis instead of mitosis. This is because we messed up while reading the script, but when Allan went back he added the corrections to the comments. When we were grading our selves, we decided that we didn't adequately describe synthesis and why it happened. Synthesis happens because for the cells to undergo mitosis and meiosis, all the chromosomes must be duplicated. We also think that we could have demonstrated that we understood our model and the process by using a thin piece of candy and using another one just like it to show that when replicating, an identical copy is created. Also, we realized that our video was too focused on mitosis and meiosis and not meiosis and fertilization. However we do believe that we explained mitosis well, but at the cost of meiosis 2 and fertilization being well explained. In general meiosis 2 could've used more explanation and we could've focused on using our model to explain it as opposed to using images because meiosis 2 slightly differs from meiosis. Along with meiosis 2 we could've improved the portrayal of fertilization. We should have modeled it using candy instead of using images as a crutch. Also we could've connected meiosis and fertilization through voice over in order to convey that they aren't really independent processes. Meiosis is the process which creates gametes, but it is also key to fertilization. When two gametes unite during fertilization, the chromosome reduction which occurred in meiosis allows the offspring contain the correct number of chromosomes. Also our model, like all others had inaccuracies For our model we used candy and attempted to use it to show the steps of meiosis but we were limited by the physical properties of the modeling material.
Dog Cloning Headlines -- Again
Published by Pete Shanks, this article titled Dog Cloning Headlines -- Again details how Barbara Streisand paid tens of thousands of dollars to have her dog cloned into two new clones. The article describes how ineffective ,and unsafe for the surrogate dog mother, this process is. 60% of the time the surrogate mother miscarries and loses the puppy. Also many of the times when these clones are born they are born with defects like “'skeletal malformations, generally not crippling though sometimes serious and always worrisome.'" The article comes to the conclusion that this process is one of animal cruelty. The article also describes how Streisand noticed the personality differences between the clones and the original. This makes me wonder, is it possible, or will it ever be possible to clone an organism to be the same physically as the original, but also be behaviorally identical? Also what causes the issues in the birth of the puppy clones?
PCR of Lambda Lab
Intro
PCR is used to replicate sections of DNA. PCR works because you only need a small sample to be able to replicated it. PCR can be used in finger printing and genome mapping. For this lab we did a PCR of Lambda DNA. The purpose of this lab was to experience PCR and continue practicing lab skills.
Procedure
On the first day me and Kellen poured the agarose gel for our group. We used all wells in the gel except for two. We used those wells for the control and standard runs. After that we prepared our reaction tubes. We used gloves so as not to contaminate the tubes. We prepared the control tube which was the same thing but was not run through the thermocycler. Each person had an individual tube which contained a small bead of TAQ polymerase and other enzymes. We labeled them in order to distinguish between them later on. When we got the tubes we held them in an ice bath and throughout the experiment we made an effort to only handle the tubes when necessary in order to keep them cool. We added 20 micro-liters of DNA primer and then 10 microliters of Lambda DNA to the tubes. After we loaded the tubes we placed them into the thermocycler and let them cycle. We then ran the gels. We took the tubes from the thermocycler and added 4 microliters of Uview to our tubes so that the DNA would be visible under UV light. We loaded our mixtures, the standard, and the control into the gel. Then we ran the gel from 9:46 AM to 10:16AM at 110V. In the middle of the run time due to time restrictions the voltage was bumped up to 120V. We loaded the gels into the UV light box to view the DNA.
Results

Conclusion
PCR or Polymerase chain reaction is a technique used to replicate a specific fragment of DNA. After the sample is run through the thermocycler we dye it, place it in a gel and electrify it to reveal the different DNA fragments. We prepared the control the same way as the other tubes but we didn't run it through the thermocycler. The control traveled 1.45 cm and the thermocycled samples traveled 4.1- 4.2 cm . Since the control had around 10,850 base pairs this shows us the importance of heat in a PCR reaction. By adding heat to a reaction it lets you isolate certain parts of the DNA. The experimental samples had roughly the same number of base pairs, which speaks to the accuracy of the PCR technique. My question is how do you know what heat to run the samples at?
Discussion
To start PCR you have to choose what DNA you want to duplicate. Afterwards add the primer and place the sample in the thermocycler. When in the thermocycler the sample is heated and cooled to denature DNA and get a specific part of the DNA to be annealed.
DNA replication duplicates DNA so that cells can continue to multiply without losing any information in replication. PCR does the same thing except for the difficulty of replication helicase enzymes and primers found in the cell. PCR uses heat to denature the DNA while in the cell a helicase enzyme is used to denature the DNA. In DNA replication an RNA primer is used, but in PCR a DNA primer is used. Polymerase can't attach to a DNA strand unless there is a primer to bind to. In the cell RNA primers have to be used because there is no excess DNA to use. The bacteria has to be used because it can survive the high temperatures.
To see DNA you have to stain it, because it's colorless naturally. The known size of the amplicon is extremely different from our results. If the known size is 1,106 base pairs and the average number our experiment showed was 284 base pairs that means there must be some kind of inaccuracies. It's possible that this was caused through hand graphing errors.
DNA replication duplicates DNA so that cells can continue to multiply without losing any information in replication. PCR does the same thing except for the difficulty of replication helicase enzymes and primers found in the cell. PCR uses heat to denature the DNA while in the cell a helicase enzyme is used to denature the DNA. In DNA replication an RNA primer is used, but in PCR a DNA primer is used. Polymerase can't attach to a DNA strand unless there is a primer to bind to. In the cell RNA primers have to be used because there is no excess DNA to use. The bacteria has to be used because it can survive the high temperatures.
To see DNA you have to stain it, because it's colorless naturally. The known size of the amplicon is extremely different from our results. If the known size is 1,106 base pairs and the average number our experiment showed was 284 base pairs that means there must be some kind of inaccuracies. It's possible that this was caused through hand graphing errors.
DNA Replication
Partners: Jackie Gibson & Kellen Mayberry
DNA replication is the process in which DNA or deoxyribonucleic acid undergoes to pass on inheritable traits or genes. DNA is the basis of all life and provides “instructions” as to what certain organisms will become and the genetic traits they will carry. DNA’s shape is vital to its functions; DNA forms in a double helix or ‘twisted ladder’ when put into layman’s terms. Each side of the double helix is composed of alternating deoxyribose and phosphate molecules. The two sides of the DNA molecule run opposite one another so that there are never two 5’ or 3’ at the same end. The 5’end is where a free phosphate group is attached to deoxyribose sugar and the 3’end is where a free hydroxyl group is attached to a deoxyribose sugar. In between the two sides of a DNA strand there are 4 different nitrogenous bases which attach the two sides together and make the DNA molecule one. The four different nitrogenous bases are: adenine (A), cytosine (C), guanine (G), and thymine (T). Adenine can only ever attach to thymine and guanine can only attach to cytosine. The bonds which hold together the sides of a DNA molecule are much stronger than the internal bonds between the nitrogenous bases. This is because the bond between the deoxyribose and phosphate molecules is a covalent phosphodiester bond and the bonds between the nucleotides are hydrogen bonds.
The first step in DNA replication begins with an enzyme known as helicase. Helicase is able to “unzip” the two sides of a DNA molecule by breaking the hydrogen bonds between each of the bases. As helicase unzips it creates a leading strand and a lagging strand. The leading strand is always of the 3’ to 5’ orientation because it is more efficient for the cell to copy this way. Once the strands are separated into leading and lagging strands RNA primase starts to prime the strands for replication. The RNA primase continuously primes the leading strand while the lagging strand gets primed in small chunks due to the differing orientations of strands. After DNA is primed DNA polymerase III attached nucleotides to already existing bases. On the lagging strand, this process of replication is not as smooth because the DNA polymerase can only copy in the 3’ to 5’ direction. Because of this the DNA polymerase works backwards and fills in the chunks that are known as Okazaki fragments. Since there is a space between fragments another enzyme is needed to help fill in the gaps, this enzyme is known as DNA ligase. The final step of DNA replication is for DNA polymerase I to go back and replace the spots on the lagging strand where the RNA primer placed uracil instead of thymine.
Bibliography
Cs.boisestate.edu, cs.boisestate.edu/~amit/teaching/342/lab/structure.html.
“The Purpose of Antiparallel DNA Strands in the DNA Molecule.” Bright Hub, 13 Nov. 2008, www.brighthub.com/science/genetics/articles/14869.aspx.
“The Purpose of Antiparallel DNA Strands in the DNA Molecule.” Bright Hub, 13 Nov. 2008, www.brighthub.com/science/genetics/articles/14869.aspx.
DNA replication is the process in which DNA or deoxyribonucleic acid undergoes to pass on inheritable traits or genes. DNA is the basis of all life and provides “instructions” as to what certain organisms will become and the genetic traits they will carry. DNA’s shape is vital to its functions; DNA forms in a double helix or ‘twisted ladder’ when put into layman’s terms. Each side of the double helix is composed of alternating deoxyribose and phosphate molecules. The two sides of the DNA molecule run opposite one another so that there are never two 5’ or 3’ at the same end. The 5’end is where a free phosphate group is attached to deoxyribose sugar and the 3’end is where a free hydroxyl group is attached to a deoxyribose sugar. In between the two sides of a DNA strand there are 4 different nitrogenous bases which attach the two sides together and make the DNA molecule one. The four different nitrogenous bases are: adenine (A), cytosine (C), guanine (G), and thymine (T). Adenine can only ever attach to thymine and guanine can only attach to cytosine. The bonds which hold together the sides of a DNA molecule are much stronger than the internal bonds between the nitrogenous bases. This is because the bond between the deoxyribose and phosphate molecules is a covalent phosphodiester bond and the bonds between the nucleotides are hydrogen bonds.
The first step in DNA replication begins with an enzyme known as helicase. Helicase is able to “unzip” the two sides of a DNA molecule by breaking the hydrogen bonds between each of the bases. As helicase unzips it creates a leading strand and a lagging strand. The leading strand is always of the 3’ to 5’ orientation because it is more efficient for the cell to copy this way. Once the strands are separated into leading and lagging strands RNA primase starts to prime the strands for replication. The RNA primase continuously primes the leading strand while the lagging strand gets primed in small chunks due to the differing orientations of strands. After DNA is primed DNA polymerase III attached nucleotides to already existing bases. On the lagging strand, this process of replication is not as smooth because the DNA polymerase can only copy in the 3’ to 5’ direction. Because of this the DNA polymerase works backwards and fills in the chunks that are known as Okazaki fragments. Since there is a space between fragments another enzyme is needed to help fill in the gaps, this enzyme is known as DNA ligase. The final step of DNA replication is for DNA polymerase I to go back and replace the spots on the lagging strand where the RNA primer placed uracil instead of thymine.
Bibliography
Cs.boisestate.edu, cs.boisestate.edu/~amit/teaching/342/lab/structure.html.
“The Purpose of Antiparallel DNA Strands in the DNA Molecule.” Bright Hub, 13 Nov. 2008, www.brighthub.com/science/genetics/articles/14869.aspx.
“The Purpose of Antiparallel DNA Strands in the DNA Molecule.” Bright Hub, 13 Nov. 2008, www.brighthub.com/science/genetics/articles/14869.aspx.
Partners : Vaishnavi Kumar, Elaine Wang
Introduction : In this lab we learned how to perform techniques needed in a DNA lab. Instead of using DNA samples, we used M&M dye to learn how we would perform labs like this. We learned how to micropipette when we measured dyes, and how to create gels and load the samples into the gel wells, and how to perform electrophoresis, which is used to separate the macromolecules according to their charge and size in the DNA samples. The molecules “migrate towards the opposite charge. A molecule with a negative charge will therefore be pulled towards the positive end” (Your Genome).
Methods : All of the group members used different colored M&M's, and we extracted the colors using a solution and put them in tubes. To make the gel, we wrapped the tray in electric gel to keep the gel from escaping the tray and to mold the gel. To create the gel Dr. Shingleton poured liquid Agarose gel into the tray, and before it solidified we placed the comb in the center of the tray to mold the wells into the gel. After we poured buffer over the gel we labeled the wells ABCD- Vaish Gray Elaine. To do the electrophoresis we covered the tray and plugged it into a power source. We ran the gel at 100V for 17 minutes.
Results : The data table shows the wells' number of dye bands, the distance each travels and the direction each travels.
Conclusion : We watched the dyes move through the gel by electrophoresis. During the lab we found that smaller fragments travel faster and further from the wells. After measuring the distance the dye traveled, we determined that my M&M dye sample traveled the furthest at 3.40cm, because it has the smallest molecule. The shortest migration distance was Vaishnavi's dye which traveled 1.70cm, which means it was the largest molecule. The agarose gel makes the smallest molecules travel faster and farther and larger molecules travel slower and shorter lengths. The molecules were negatively charged and traveled towards the positive end of the gel. Blue, yellow, and orange dyes were found in our M&M samples which is logical because Vaish had a green M&M, I had a yellow M&M, and Elaine had an orange M&M. The blue dye traveled the shortest distance and was the largest molecule and was extremely concentrated the samples that it was in. The yellow dye traveled the farthest, meaning it was the smallest molecule. The third lane, where we loaded “sample C,” had the most variation and the largest amount of bands. The three bands it dispersed into were the colors blue, red and yellow. The other lanes only had one and two bands. The orange and pinkish dyes went the same distance which shows that the molecules might be similar. Our experiment went well, and our results are logical results in my opinion.
Discussion : Size is key in analyzing DNA. Smaller fragments of DNA travel faster inagarose gel, while larger fragments travel slower. DNA molecules with differing weights like a sample that is 700 daltons would travel faster than a sample that is 4000 daltons. The different bands represent certain amounts of base pairs. The bands closer to the starting lanes contain the higher amount of negative base pairs, and the bands further away away contain the positive base pairs.
In DNA the sugar phosphate backbone is negatively charged. That's why DNA travels in a positive direction in the gel because in electrophoresis one side of the gel has a negative charge and the other a positive charge. We didn't actually use DNA but our molecules were negatively charged so we would be able to observe movement like with DNA. Fast Green FCF would be the best to use in this lab because it has the most negative formal charge. Although Betanin has a negative charge on the oxygen, the positive charge on the nitrogen cancels it out, giving it an overall formal charge of zero. Carminic acid and citrus red 2 won't function because they're chargeless, meaning it would lack attraction towards either side of the gel.
The pore size of the gel is important to validating the data. The gel’s concentration determines how DNA (or other sample) travels. A higher concentration gel would cause DNA fragments to not travel very hard because of its smaller pore size. A gel with a lower concentration would facilitate easier DNA travel because the pore sizes are bigger. Each have their benefits and pitfalls. A high concentration are more appropriate for smaller sized DNA because they can contain the samples better because the smaller DNA run more. For larger DNA it would be more difficult to see. For a lower concentration, it would be better for larger sized DNA, since larger fragments travel slower and this would allow an even variation among the gel rather than it being compacted. It would not be ideal to use low concentration on smaller sized DNA in fear of it running off the gel.
In DNA the sugar phosphate backbone is negatively charged. That's why DNA travels in a positive direction in the gel because in electrophoresis one side of the gel has a negative charge and the other a positive charge. We didn't actually use DNA but our molecules were negatively charged so we would be able to observe movement like with DNA. Fast Green FCF would be the best to use in this lab because it has the most negative formal charge. Although Betanin has a negative charge on the oxygen, the positive charge on the nitrogen cancels it out, giving it an overall formal charge of zero. Carminic acid and citrus red 2 won't function because they're chargeless, meaning it would lack attraction towards either side of the gel.
The pore size of the gel is important to validating the data. The gel’s concentration determines how DNA (or other sample) travels. A higher concentration gel would cause DNA fragments to not travel very hard because of its smaller pore size. A gel with a lower concentration would facilitate easier DNA travel because the pore sizes are bigger. Each have their benefits and pitfalls. A high concentration are more appropriate for smaller sized DNA because they can contain the samples better because the smaller DNA run more. For larger DNA it would be more difficult to see. For a lower concentration, it would be better for larger sized DNA, since larger fragments travel slower and this would allow an even variation among the gel rather than it being compacted. It would not be ideal to use low concentration on smaller sized DNA in fear of it running off the gel.
Works Cited :
“What Is Gel Electrophoresis?” Facts, The Public Engagement Team at the Wellcome Genome Campus, 25 Jan. 2016
“Gel Electrophoresis (Article).” Khan Academy
“What Is the Pore Size, Porosity and Pore Size Distribution inside Agarose Gel Used in Gel Electrophoresis?” Quora, Umer Bakali
“Molecular Facts and Figures.” Integrated DNA Technologies
“What Is Gel Electrophoresis?” Facts, The Public Engagement Team at the Wellcome Genome Campus, 25 Jan. 2016
“Gel Electrophoresis (Article).” Khan Academy
“What Is the Pore Size, Porosity and Pore Size Distribution inside Agarose Gel Used in Gel Electrophoresis?” Quora, Umer Bakali
“Molecular Facts and Figures.” Integrated DNA Technologies